We are happy to announce that a new SINGEK paper is out in Open Biology:
G. Leonard, A. Labarre, D. S. Milner, A. Monier, D. Soanes, J. G. Wideman, F. Maguire, S. Stevens, D. Sain, X. Grau-Bové, A. Sebé-Pedrós, J. E. Stajich, K. Paszkiewicz, M. W. Brown, N. Hall, B. Wickstead and T. A. Richards
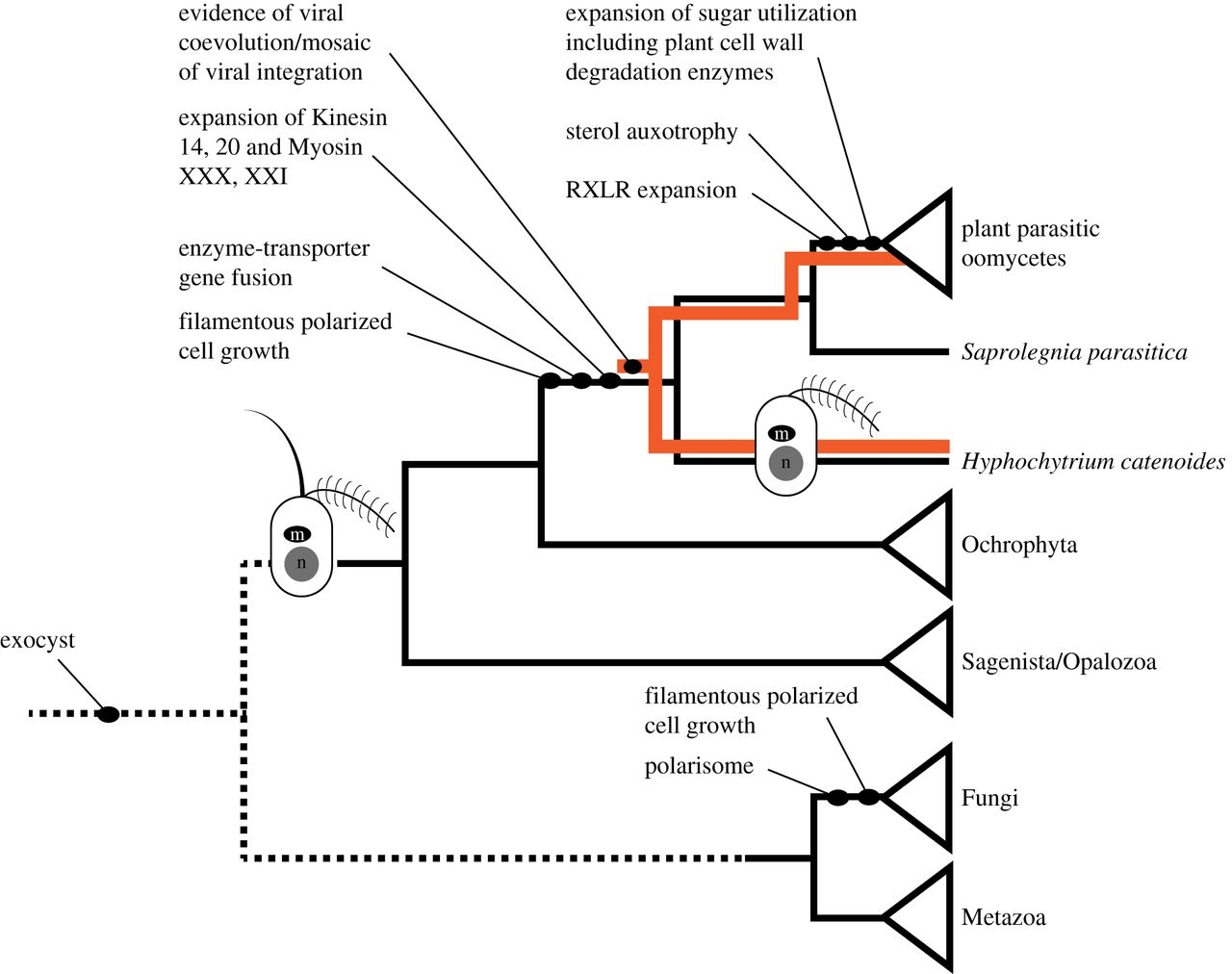
Schematic phylogenetic tree summarizing the evolution of cell and genomic characters relevant to the evolution of the Pseudofungi.
Eukaryotic microbes have three primary mechanisms for obtaining nutrients and energy: phagotrophy, photosynthesis and osmotrophy. Traits associated with the latter two functions arose independently multiple times in the eukaryotes. The Fungi successfully coupled osmotrophy with filamentous growth, and similar traits are also manifested in the Pseudofungi (oomycetes and hyphochytriomycetes). Both the Fungi and the Pseudofungi encompass a diversity of plant and animal parasites. Genome-sequencing efforts have focused on host-associated microbes (mutualistic symbionts or parasites), providing limited comparisons with free-living relatives. Here we report the first draft genome sequence of a hyphochytriomycete ‘pseudofungus’; Hyphochytrium
catenoides. Using phylogenomic approaches, we identify genes of recent viral ancestry, with related viral derived genes also present on the genomes of oomycetes, suggesting a complex history of viral coevolution and integration across the Pseudofungi. H. catenoides has a complex life cycle involving diverse filamentous structures and a flagellated zoospore with a single anterior tinselate flagellum. We use genome comparisons, drug sensitivity analysis and high-throughput culture arrays to investigate the ancestry of oomycete/pseudofungal characteristics, demonstrating that many of the genetic features associated with parasitic traits evolved specifically within the oomycete radiation. Comparative genomics also identified differences in the repertoire of genes associated with filamentous growth between the Fungi and the Pseudofungi, including differences in vesicle trafficking systems, cell-wall synthesis pathways and motor protein repertoire, demonstrating that unique cellular systems underpinned the convergent evolution of filamentous osmotrophic growth in these two eukaryotic groups.











